 PSD should also be calculated for a de-trended time series. append (, ignore_index = True ) return accuracy_df def plot_pgram ( series, diff_order ): """ This function plots thd Power Spectral Density of a de-trended series. array ( y1 ))) * 100, 1 ) accuracy_df = accuracy_df. round ( rmse ( y1, y2 ), 1 ) map_error = np. around ( resid_mean, 2 )) print ( " \n ** Ljung Box Test, p-value:", np. tight_layout () print ( "** Mean of the residuals: ", np. plot ( ax = ts_ax ) plot_acf ( residuals, lags = lags, ax = acf_ax ) sns. subplot2grid ( layout, ( 1, 1 )) residuals. subplot2grid ( layout, ( 1, 0 )) kde_ax = plt. subplot2grid ( layout, ( 0, 0 ), colspan = 2 ) acf_ax = plt. figure ( figsize = ( 10, 8 )) layout = ( 2, 2 ) ts_ax = plt. mean ( ljung ( x = residuals, lags = lags )) norm_p_val = jb ( residuals ) adfuller_p = adfuller ( residuals ) fig = plt. Lags should be min(2*seasonal_period, T/5) plots from: """ resid_mean = np. If Ljung Box test shows p> 0.05, the residuals as a group are white noise. Consider different model and/or add exogenous variable. If the first two are not met, we have not fully captured the information from the data for prediction. 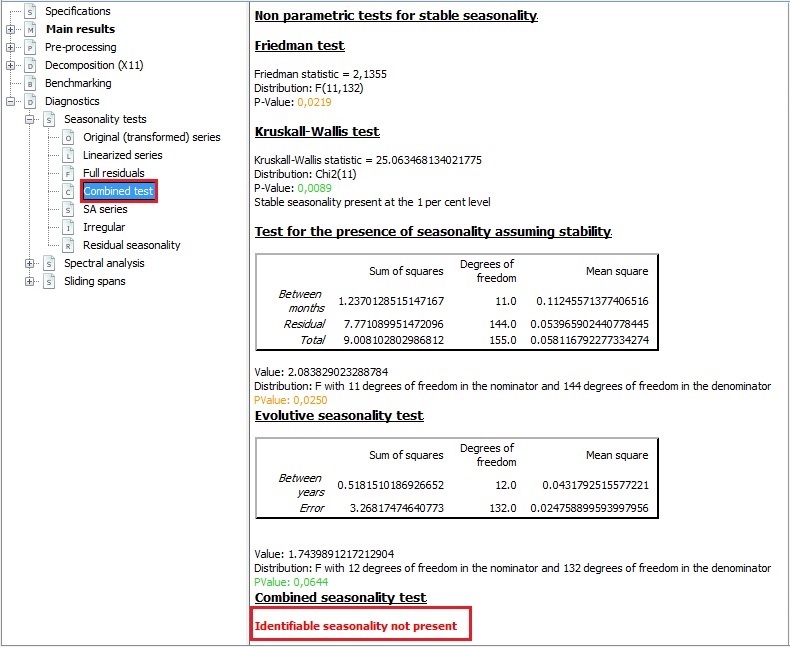 
First two are must, while last two are good to have. Ideally the residuals should be uncorrelated, zero mean, constant variance and normally distributed. abs (( y_true - y_pred ) / y_true )) * 100 def residcheck ( residuals, lags ): """ Function to check if the residuals are white noise. If the series has outliers, compare/select model using MAPE instead of rmse() """ y_true, y_pred = np. #collapse-hide # Define some custom functions to help the analysis def MAPE ( y_true, y_pred ): """ %Error compares true value with predicted value. #statistics libraries import statsmodels.api as sm import scipy from scipy.stats import anderson from _measures import rmse from import adfuller from import month_plot, seasonal_plot, plot_acf, plot_pacf, quarter_plot from import seasonal_decompose from import ExponentialSmoothing, SimpleExpSmoothing from import acorr_ljungbox as ljung #from nimbusml.timeseries import SsaForecaster from import diff as diff import pmdarima as pm from pmdarima import ARIMA, auto_arima from scipy import signal from scipy.stats import shapiro from scipy.stats import boxcox from sklearn.preprocessing import StandardScaler #library to use R in Python import rpy2 from rpy2.robjects import pandas2ri pandas2ri. 
use ( 'seaborn-white' ) % matplotlib inline #collapse-hide #Author: Sandeep Pawar #Version: 1.0 #Date import pandas as pd import numpy as np import itertools #Plotting libraries import matplotlib.pyplot as plt import seaborn as sns import altair as alt plt. 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
